Interpolation Methods
Interpolation估计是一个过程吗values that lie between known data points.
Interpolation involves the construction of a functionfthat matches givendata values,yi, at givendatasites,xi, in the sense thatf(xi) =yi, alli.
The interpolant,f, is usually constructed as the unique function of the form
that matches the given data, with the functionsfjchosen “appropriately”.
In spline interpolation, one chooses thefjto be thenconsecutive B-splinesBj(x) =B(x|tj,...,tj+k),j= 1:n, of orderkfor some knot sequencet1≤t2≤ ... ≤tn+k.
About Interpolation Methods
Method |
Description |
|---|---|
Linear |
Linear interpolation. This method fits a different linear polynomial between each pair of data points for curves, or between sets of three points for surfaces. |
Nearest neighbor |
Nearest neighbor interpolation. This method sets the value of an interpolated point to the value of the nearest data point. Therefore, this method does not generate any new data points. |
Cubic spline |
Cubic spline interpolation. This method fits a different cubic polynomial between each pair of data points for curves, or between sets of three points for surfaces. |
Shape-preserving |
Piecewise cubic Hermite interpolation (PCHIP). This method preserves monotonicity and the shape of the data. For curves only. |
Biharmonic (v4) |
MATLAB®4 For surfaces only. |
Thin-plate spline |
Thin-plate spline interpolation. This method fits smooth surfaces that also extrapolate well. For surfaces only. |
For surfaces, the Interpolant fit type uses the MATLABscatteredInterpolantfunction for linear and nearest methods, and the MATLABgriddatafunction for cubic and biharmonic methods. The thin-plate spline method uses thetpapsfunction.
interpolant使用的类型取决于characteristics of the data being fit, the required smoothness of the curve, speed considerations, post-fit analysis requirements, and so on. The linear and nearest neighbor methods are fast, but the resulting curves are not very smooth. The cubic spline and shape-preserving and v4 methods are slower, but the resulting curves are very smooth.
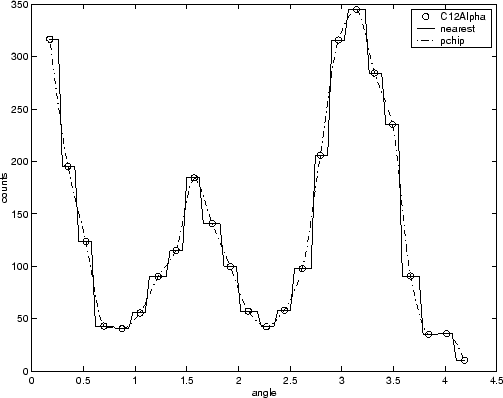
For example, the nuclear reaction data from thecarbon12alpha.matfile is shown here with a nearest neighbor interpolant fit and a shape-preserving (PCHIP) interpolant fit. Clearly, the nearest neighbor interpolant does not follow the data as well as the shape-preserving interpolant. The difference between these two fits can be important if you are interpolating. However, if you want to integrate the data to get a sense of the total strength of the reaction, then both fits provide nearly identical answers for reasonable integration bin widths.
Note
Goodness-of-fit statistics, prediction bounds, and weights are not defined for interpolants. Additionally, the fit residuals are always 0 (within computer precision) because interpolants pass through the data points.
Interpolants are defined aspiecewise polynomialsbecause the fitted curve is constructed from many “pieces” (except forBiharmonicfor surfaces which is a radial basis function interpolant). For cubic spline and PCHIP interpolation, each piece is described by four coefficients, which the toolbox calculates using a cubic (third-degree) polynomial.
Refer to the
splinefunction for more information about cubic spline interpolation.Refer to the
pchipfunction for more information about shape-preserving interpolation, and for a comparison of the two methods.Refer to the
scatteredInterpolant,griddata, andtpapsfunctions for more information about surface interpolation.
It is possible to fit a single “global” polynomial interpolant to data, with a degree one less than the number of data points. However, such a fit can have wildly erratic behavior between data points. In contrast, the piecewise polynomials described here always produce a well-behaved fit, so they are more flexible than parametric polynomials and can be effectively used for a wider range of data sets.

